New Projects Sprouting: Draba and the Beech Family
Hello friends,
Two new workflows are sprouting up on Notes from Nature: Capture the Collections – and we think you’ll go nuts for them!
 All the Draba: Tales of Whitlow grasses
All the Draba: Tales of Whitlow grasses
Small but mighty, Draba belong to the mustard family (Brassicaceae) and thrive in cold, rocky, and windswept habitats. Sometimes called “Whitlow grasses,” these hardy wildflowers are found across North America and around the world, from Arctic tundra to high mountain slopes. Draba often grow in disturbed areas – roadsides, pastures, and gravelly soils – and are among the first plants to bloom in spring, covering the ground with tiny white or yellow flowers. Though small, Draba play a big role in helping scientists understand how plants adapt and survive in extreme environments.
 Notes from Nuts
Notes from Nuts
Now for something a little nutty: the Fagaceae – also known as the Beech family – include some very iconic trees: oaks, beeches, and chestnuts. These trees are nut-orious! They form the backbone of many forests across the Northern Hemisphere, stabilizing soils, sheltering wildlife, and producing the acorns and nuts that sustain entire ecosystems.
Wood from these trees has shaped cultures and economies for centuries – from furniture and flooring to barrels and instruments. But their scientific value is just as enduring: studying these trees helps us understand forest health, climate resilience, and long-term ecological change.
As always, your contributions make these collections searchable, shareable, and scientifically meaningful. Whether you’re here for the alpine flowers or the mighty nut – bearing trees, join us in exploring biodiversity one label at a time.
– Kim, Lyn, and Jennifer – The Capture the Collections Team


Nouveaux projets en germination : Draba et la famille des hêtres
Bonjour les ami·e·s,
Deux nouveaux flux de travail bourgeonnent sur Notes de la nature : Capturer les collections – et on pense que vous allez en devenir fous de nature!
 Tous les Draba : histoires de drabes
Tous les Draba : histoires de drabes
Petits mais puissants, les Draba font partie de la famille de la moutarde (Brassicaceae) et prospèrent dans les milieux froids, rocheux et exposés au vent. Parfois appelées « drabes » ou « whitlow-grasses », ces petites plantes robustes se retrouvent un peu partout en Amérique du Nord et ailleurs dans le monde, des toundras arctiques aux pentes montagneuses les plus élevées.
Les Draba poussent souvent dans les zones perturbées – le long des routes, dans les pâturages ou les sols graveleux – et comptent parmi les premières fleurs du printemps, recouvrant le sol de minuscules fleurs blanches ou jaunes. Malgré leur taille, elles jouent un grand rôle pour aider les scientifiques à comprendre comment les plantes s’adaptent et survivent dans des environnements extrêmes.
 Notes du noyer
Notes du noyer
Passons maintenant à quelque chose d’un peu plus casse-noisette! Les Fagaceae – aussi connues sous le nom de famille des hêtres – comprennent des arbres emblématiques comme les chênes, les hêtres et les châtaigniers. Ces arbres sont de véritables vedettes de la forêt : ils forment l’épine dorsale de nombreux écosystèmes à travers l’hémisphère Nord, stabilisent les sols, abritent la faune et produisent les glands et les noix qui nourrissent des écosystèmes entiers.
Leur bois a façonné des cultures et des économies depuis des siècles – meubles, planchers, tonneaux et instruments de musique – et leur valeur scientifique demeure tout aussi durable. Étudier ces arbres nous aide à mieux comprendre la santé des forêts, leur résilience face au climat et les changements écologiques à long terme.
Comme toujours, vos contributions rendent ces collections consultables, partageables et scientifiquement précieuses. Que vous soyez là pour les fleurs alpines ou les arbres à noix majestueux, joignez-vous à nous pour explorer la biodiversité, une étiquette à la fois.
— Kim, Lyn et Jennifer – L’équipe de Capturer les collections
WeDigBio – One week away!
WeDigBio starts in less than a week. The event takes place October 9-12, 2025. People from all over the world join together to digitize museum specimen data and to celebrate biodiversity collections. We hope you will join us at Notes from Nature.
Notes from Nature will have lots of great expeditions to contribute to. We’ll be featuring California plants, bees, environmental archaeology specimens and more, so please stop by. It’s also very helpful if you can spread the word about this unique and important event.
— The Notes from Nature Team
Introducing ArchaeoZooArchive – An Archaeological Natural History Archive!
The Florida Museum Environmental Archaeology Program is excited to bring a new adventure to all you Notes from Nature fans … the ArchaeoZooArchive … the first archaeological natural history transcribing project in Notes from Nature!
When we ask you, the wonderful community scientists, for help transcribing nature from natural history collections, we are typically talking about data from plant and animal specimens that have been collected over the past 100 years or so – maybe even as far back as 200 years! These records provide excellent insight to our modern biodiversity and environments and the data you transcribe is vital for understanding how we should cherish and manage our natural world. But you might be interested to know that all that modern specimen data provides only a TINY glimpse of a very long history of people’s relationships with nature. When you consider that people have been admiring, using, managing, and impacting natural resources for over a million years, you realize that 200 years is but the blink of an eye. So we bring you an opportunity to help scientists look deeper into time …
Here at the Florida Museum of Natural History, the Environmental Archaeology Program is devoted to the study of the deep history of human-environment interactions using the remains of plants, animals, and landscapes from archaeological sites. The Environmental Archaeology collections include literally millions of specimens from more than 800 archaeological sites representing over 10,000 years of prehistory across the circum-Caribbean (the region that includes the SE USA, Central America, Caribbean, and northern South America). With assistance from the National Science Foundation (NSF Grant DBI #1929448) we have been working hard to make these specimens more accessible to scientists and the community through improvements in curation, archival documentation, and of course, making the data about these specimens available through both specialized and open data publishers. The ArchaeoZooArchive is one step in the process – inviting the community to help us turn our data records about animal remains from archaeological sites into usable data, one mysterious specimen card at a time!
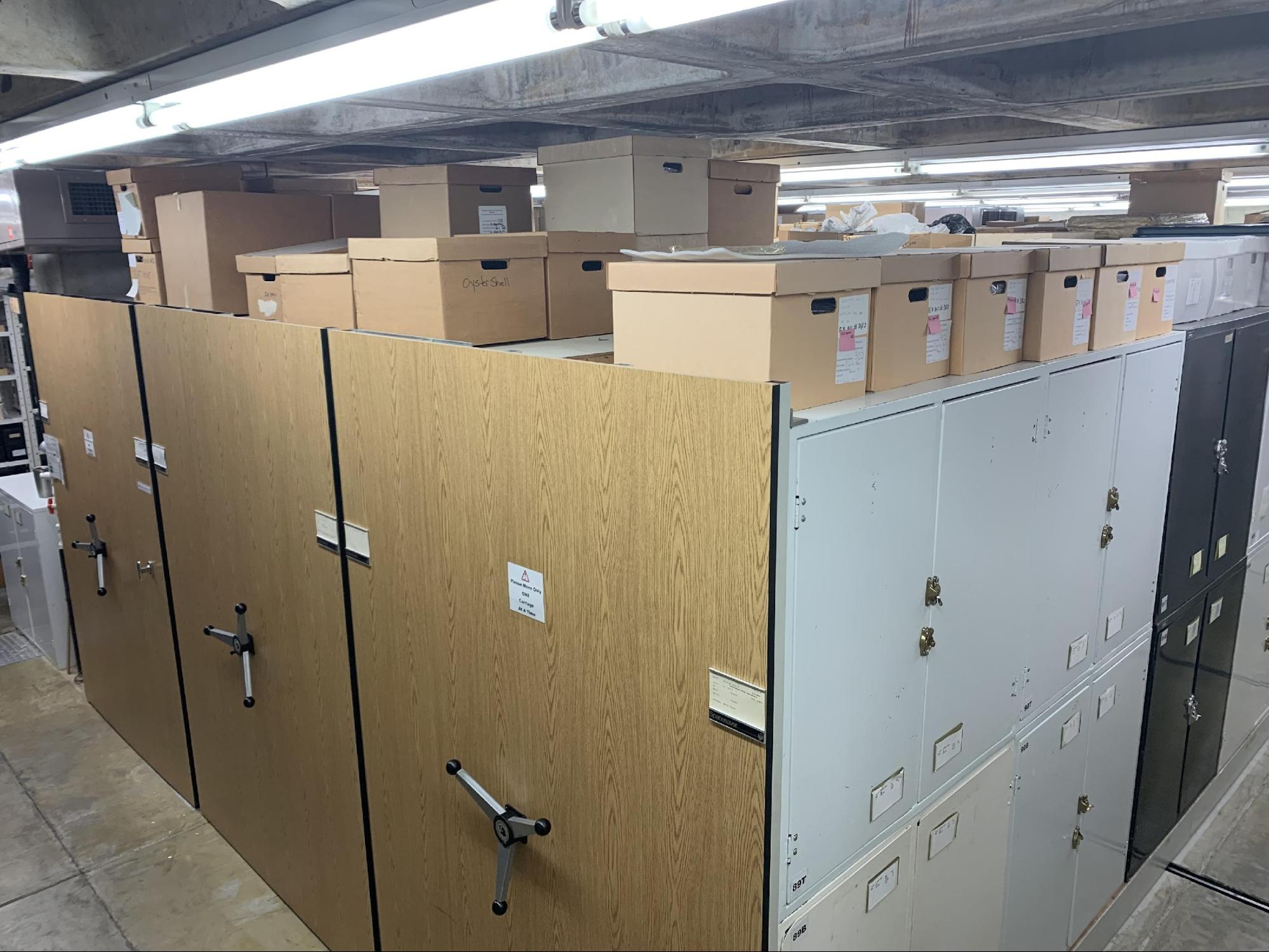



The data collected from the EAP specimens has informed us about the longer history of how environments have impacted people and how, in turn, people have impacted their environments. But surprisingly little of our data has been fully digitized, so although much has already been learned, by making our data available more broadly, we can learn even more valuable lessons relevant to today’s major issues by documenting the millennia of “human experiments” of the past – learning from past successes and mistakes.
Those of you who have been actively engaged with community science know this part of the story all too well – alongside the vast EAP collections are the MILLIONS of associated data records – as accession folders, card cabinets, and notebooks …



Over the years we have made great strides in scanning our primarily paper records into digital formats that can be more easily shared.

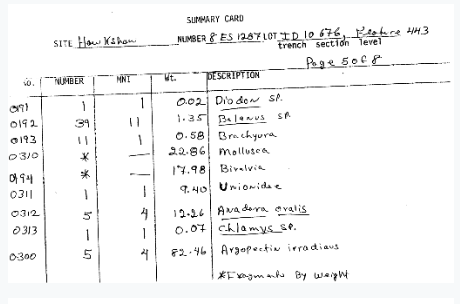
But even with digital versions, it is an archaeological project of its own to dig through these and organize, standardize, and make available the actual specimen-by-specimen data in these records.
This is where Notes from Nature has provided a wonderful opportunity to reach out to community scientists and volunteers to help us digitize the data from our scanned paper records. In collaboration with our lab team of students, researchers, and volunteers, and with the special assistance of a dedicated teen volunteer from a local high school (stay tuned to hear more from them in our next blog post), and of course with Mike Denslow of Notes from Nature, we have created a first expedition to digitize our most straight-forward data records, the Environmental Archaeology zooarchaeological specimen data summary cards! We started with zooarchaeology because it has historically been the primary focus of our collections and thus is the collection with the most standardized data recording methods and broadly accepted taxonomies in our archives. But don’t let this fool you – it will still be a challenge that we hope you’ll be excited to take on!
Today we launched ArchaeoZooArchive expeditions by geographic region, starting with South American collections! Once those have been completed we will move on to Central American and Caribbean expeditions.
Please give ArchaeoZooArchive: The South American Expedition
a try today and let us know what you think.
— Kitty F. Emery, Jessica Nickles King, Nicole Fuller, and Al Keller.
EAP logo by Al Keller. Other images by EAP.
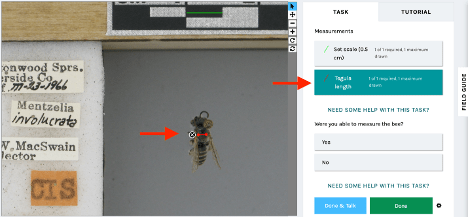
Measuring bees for Climate Change: California Edition
We are excited to launch a new measurement expedition, Measuring bees for Climate Change: California Edition. This project brings together specimen images from six institutions across the US, featuring California bees from the past 100 years. These measurements will contribute to a new study aimed at answering the question: is climate change shrinking bees?
Recently, scientists have documented body size declines in animal groups over the past 50-100 years. For insects, the causes of this shrinking effect are strongly debated. One possibility is warming temperatures interfering with normal animal development. Climate change may also affect size by restricting food availability in some environments. Regardless of cause, changes in body size can have major ecological consequences. Smaller bees, for example, carry less pollen and may be more sensitive to environmental disturbances, restricting their ability to deliver critical pollination services.


California is home to approximately 1,600 bee species, thanks to its diverse range of habitats, which include deserts, forests, and grasslands. California’s climate is warming and becoming more unpredictable, with complex and unknown impacts on bees. Notes from Nature volunteers can help us understand body size changes in California’s bees by capturing critical measurement data on historical specimens . You’ll help us measure bee body width (or intertegular distance–essentially, shoulder width), which provides a good estimate of overall body size.
Thanks to existing Notes from Nature volunteer effort, we were able to show the value of citizen community science approaches to bee measurements. In a recent paper we found that Notes from Nature volunteers provide highly accurate bee body size measurements, differing only 2% from trained researchers (link to study). This study showed the enormous potential for community scientists to unlock the wealth of size data stored in museum records.
Our preliminary results hint at exciting potential links between bee size and climate–more measurements will help fill in some critical gaps that we haven’t been able to cover yet. We need help completing these measurements to expand our dataset to represent a greater diversity of bees over time and in different habitats. This data will help us answer questions like: are bees really shrinking? Are males and females shrinking at different rates? Do hotter regions cause greater shrinking? Which habitats are most vulnerable to these effects? Questions like this require a huge amount of data and have not yet been answered for bees at this scale. Community science measurements will help us overcome this data challenge and provide new insights into bee biology in a warming world.
Please visit Notes from Nature – Big Bee Bonanza! today to help us complete the Measuring bees for Climate Change: California Edition expedition.
Notes from Nature: Capture the Collections updates
Hello friends,
Notes from Nature: Capture the Collections is thrilled to share some updates with you!
Maple Nation: Acers of Canada
We’ve just launched a brand-new workflow focused on Canada’s iconic maple trees (Acer). From the Sugar Maple (Acer saccharum) behind maple syrup to the towering Bigleaf Maple (Acer macrophyllum) spreading its canopy along the Pacific coast, these trees are central to our landscapes, culture, and history. In this workflow, you’ll be helping us transcribe collector information, locality details, and coordinate values.
We continue to love the OAC Herbarium!
Thanks to your help, the fern workflow at the University of Guelph’s OAC Herbarium was a great success! The project now continues with another exciting group: the Asteraceae, also known as the daisy or sunflower family. Among the most diverse plant families worldwide, they occur in nearly every Canadian ecosystem, including the Artic, to the Rockies, boreal forests, prairies and coastal sand dunes. They have plenty of stories to tell, and your transcriptions will help bring those stories to light. In this workflow, you’ll be capturing collector details (name, date, collector number) to make these specimens more accessible.
As always, the label data you capture supports scientists, historians, educators, and conservationists. Your contributions make these collections searchable, shareable, and meaningful for people everywhere.
Thank you for being part of Capture the Collections—we couldn’t do this without you! And don’t forget to share your discoveries on the Talk boards.
Kim, Lyn and Jennifer
The Capture the Collections team
Start transcribing on Zooniverse
Learn more about the project here
Bonjour les amis
Notes de la nature – Capturer des collections est ravis de partager quelques nouveautés avec vous!
La nation de l’érable : Acers du Canada
Nous venons de lancer un tout nouveau flux de travail consacré aux emblématiques érables du Canada (Acer). De l’Érable à sucre (Acer saccharum), qui nous donne le sirop d’érable, à l’imposant Érable à grandes feuilles (Acer macrophyllum), qui étend sa canopée le long de la côte Pacifique, ces arbres occupent une place centrale dans nos paysages, notre culture et notre histoire. Dans ce flux de travail, vous nous aiderez à transcrire les informations sur les collecteurs, les données de localité et les coordonnées.
Nous continuons d’adorer l’herbier OAC!
Grâce à votre aide, le flux de travail sur les fougères de l’herbier OAC de l’Université de Guelph a été un grand succès! Le projet se poursuit maintenant avec un autre groupe passionnant : les Asteraceae, aussi appelées la famille des marguerites ou des tournesols. Parmi les familles de plantes les plus diversifiées au monde, on peut les retrouver dans quasi tous les habitats et écosystèmes canadiens, incluant l’Arctique, les Rocheuses, la foret boréale, les praires et les dunes côtières. eElles ont beaucoup d’histoires à raconter -et vos transcriptions contribueront à les mettre en lumière. Dans ce flux de travail, vous saisirez les détails sur les collecteurs (nom, date, numéro du collecteur) afin de rendre ces spécimens plus accessibles.
Comme toujours, les données d’étiquettes que vous capturez soutiennent les scientifiques, les historiens, les éducateurs et les spécialistes de la conservation. Vos contributions rendent ces collections consultables, partageables et utiles pour tous.
Merci de faire partie de Capture the Collections — nous ne pourrions pas y arriver sans vous! Et n’oubliez pas de partager vos découvertes sur les forums de discussion.
Kim, Lyn et Jennifer
L’équipe Capture the Collections
Spotlight on the Guelph OAC Herbarium: Ferns in Focus!
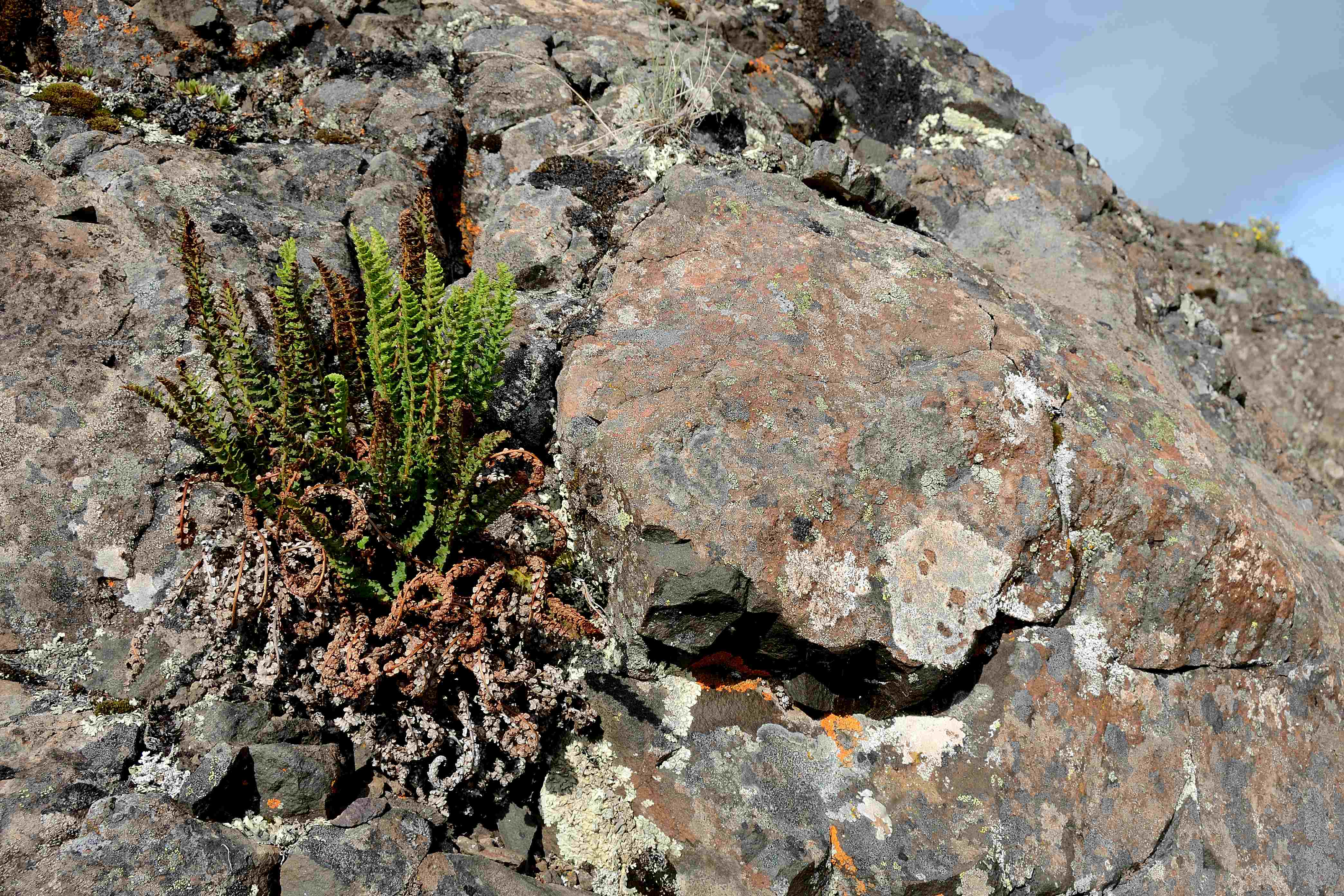


Notes from Nature: Capture the Collections is excited to be teaming up with the University of Guelph’s OAC Herbarium to transcribe part of their digitized collection—starting with ferns!
The OAC Herbarium imaged most of its vascular plant specimens in 2015 through Canadensys, but many of the label details were never transcribed. To start, we’re running one new workflow focused on 1,001 fern specimens from the families:
Blechnaceae – chain ferns, hard ferns
Davalliaceae – hare’s foot ferns
Polypodiaceae – polypody and resurrection ferns
You’ll be helping us capture Collector information, making these specimens easier to search and use for research, education, and preservation of collections.
This fern collection is closely tied to the work of Professor Donald M. Britton (1923–2012), who spent his career at the University of Guelph and was considered one of North America’s top fern experts. He co-authored Ferns and Fern Allies of Canada, which is still used today.
Thanks for helping us bring more of these specimens online!
Start transcribing on Zooniverse
Learn more about this new partnership here
Pleins feux sur l’herbier OAC de Guelph : les fougères à l’honneur !
Notes de la nature – Capturer des collections est ravi de s’associer à l’herbier OAC de l’Université de Guelph pour transcrire une partie de leur collection numérisée, en commençant par les fougères !
L’herbier OAC a numérisé la plupart de ses spécimens de plantes vasculaires en 2015 grâce à Canadensys, mais de nombreux détails trouv é s sur les étiquettes n’ont jamais été transcrits. Pour commencer, nous mettons en place un nouveau processus axé sur 1 001 spécimens de fougères appartenant aux familles suivantes :
Blechnaceae – fougères en chaîne, fougères dures
Davalliaceae – fougères pied-de-lièvre
Polypodiaceae – polypodies et fougères résurrection
Vous nous aiderez à saisir les informations relatives aux collectionneurs, ce qui facilitera la recherche et l’utilisation de ces spécimens à des fins de recherche, d’éducation et de conservation des collections.
Cette collection de fougères est étroitement liée au travail du professeur Donald M. Britton (1923-2012), qui a passé sa carrière à l’université de Guelph et était considéré comme l’un des plus grands experts en fougères d’Amérique du Nord. Il est coauteur de Ferns and Fern Allies of Canada, un ouvrage qui est encore utilisé aujourd’hui.
Merci de nous aider à mettre en ligne davantage de ces spécimens !
Commencez à transcrire sur Zooniverse
Pour en savoir plus sur ce nouveau partenariat, cliquez ici.
12 years of Notes from Nature
Happy Anniversary, Happy Earth Day and most of all Thank You!
Today is the Notes from Nature projects 12th anniversary! We first launched Notes from Nature on Earth Day 2013. We are beyond grateful to still be here today and to be continuing this important work with you all every single day.
We are also thankful for all the support of our colleagues, the National Science Foundation and the Zooniverse team!
Here’s to 12 more years! As always there is more work to be done, so please stop by Notes from Nature when you can and help us spread the word about the importance of biodiversity data.
– The Notes from Nature Team
WeDigBio 2025 – Appreciation
We closed out the last day of WeDigBio April 2023 with over 2,700 classifications. That puts Notes from Nature at over 12,200 for the event. Thanks to all that made it possible!
Digitizing Biological Collections in Canada was particularly popular this time around with 4,896 classifications!
There are still lots of expeditions from plants to salamanders, bees and more available on our site. As always feel free to stop anytime and help by spreading the word about the importance of open data about biodiversity.
— The Notes from Nature Team
California Central Valley Botany
Our new expedition at #PlantstoPixels highlights the botany of the California Central Valley. Nestled between the Sierra Nevada and California Coastal Ranges, few areas in California have undergone more manmade change than the Central Valley. Before widespread agriculture, the Central Valley was home to vast wetlands and grasslands that supported fauna such as pronghorn antelope, elk, and migratory birds. The Central Valley was also once home to Tulare Lake, the largest lake by area west of the Mississippi River.
During the 20th century, vast irrigation projects combined with an explosion in agricultural output caused a sharp increase in the use of groundwater and natural water sources in the area, leading to the disappearance of marshes and lakes and subsidence due to groundwater depletion. The Central Valley is estimated to be about 28 feet lower in elevation on average than it was in 1850 due to aquifer usage. Today, the Central Valley produces one-quarter of all food in the United States and 8% of the total agricultural value, making it one of the most productive agricultural areas in the country.
Despite these dramatic changes to the Central Valley’s natural landscape, there is still a significant role that botany can play in the region. There are still remaining areas of inland chaparral, woodland, and marsh that need study and preservation. Historic botany collections also play an important role in documenting the changes the Central Valley has undergone throughout the centuries, including climate change, habitat destruction, droughts, and the introduction of invasive species.
Be sure to check out our California Central Valley Botany expedition!
Here are some native plants you may encounter while transcribing collections from the California Central Valley:
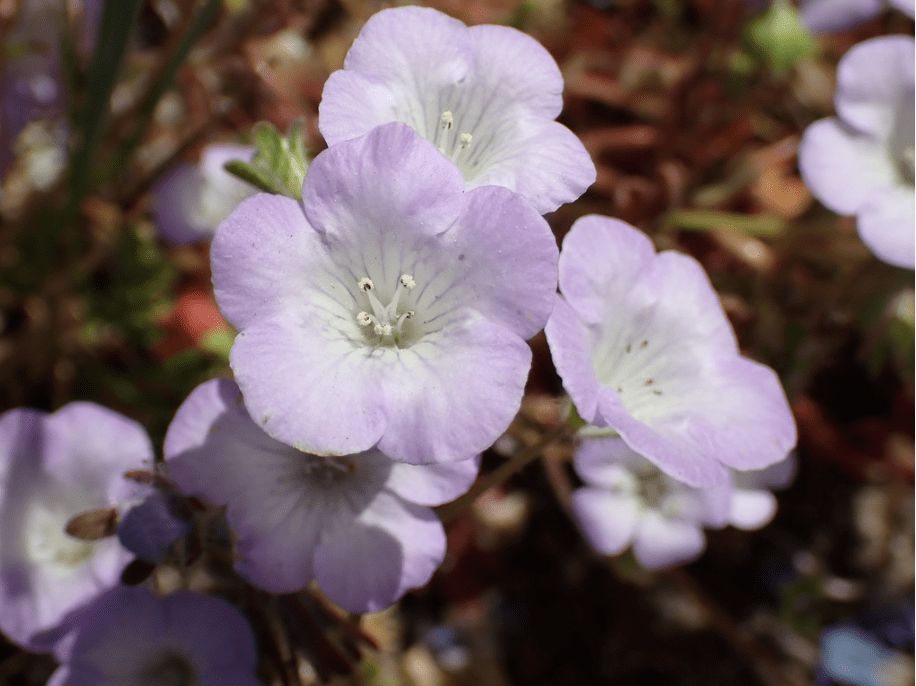
Phacelia douglasii: Phacelia douglasii or Douglas’s Phacelia is native only to California where it grows mostly in the central valley, California coastal mountain ranges and northern Mojave desert. It is a member of the Hydrophyllaceae Family, a subfamily of Boraginaceae.
Epilobium obcordatum:
Epilobium obcordatum or the Rockfringe Willowherb lives up to its name by growing in mats along rocky outcrops , fringing cracks in rocks. Although not present in the central valley itself, it can be found in the Sierra Nevadas and Sierra Nevada foothills as they rise out of the valley. This plant is in the family Onagraceae and can be found in several western states including Nevada, Idaho and California.

Linanthus pungens:
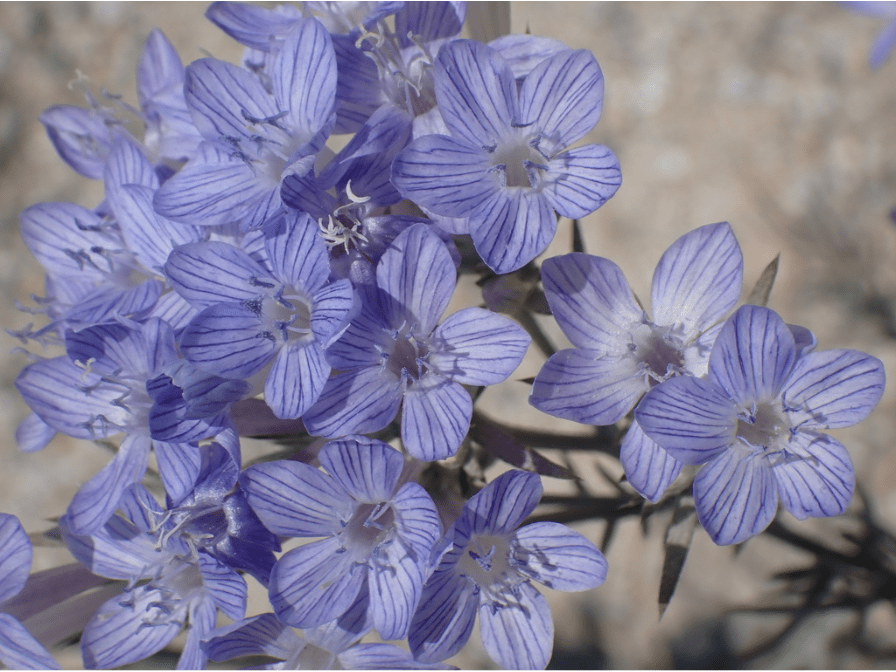
A member of the family Polemoniaceae Linanthus pungens or the Granite gilia is native to California and west coast North America. It is a very resilient plant, able to grow in desert washes ,grassland, pine forests as well as both high and low altitude environments. This plant is very dominant in El Dorado County for example, where it thrives in white pine forests of the Sierra Nevada foothills, although it can also be found in other environments throughout the central valley and surrounding areas.

Coastal California: Monterey to Santa Barbara
Our new expedition at #PlantstoPixels, explores the rich botany of California’s central coast—a region known for its stunning natural beauty and remarkable biodiversity. From towering redwood forests and dramatic sea cliffs to coastal salt marshes, this landscape supports an impressive array of plant life. This area also serves as a transition zone for marine life, sitting at the confluence of major ocean currents.
Beyond its ecological significance, the region is steeped in California’s cultural history and landmarks. South of Monterey Bay, the Santa Lucia Mountains rise to form the dramatic coastline of Big Sur. Inland, you’ll find the Salinas Valley—one of the most fertile and productive agricultural regions in the United State. The Salinas Valley and Monterey Bay region inspired many of John Steinbeck’s classic novels. Continuing south on Highway 1, you’ll pass Hearst Castle, the opulent former estate of William Randolph Hearst, before arriving in Morro Bay and San Luis Obispo. Further along the coast, in Santa Barbara, nestled against the Santa Ynez Mountains, you can see Chumash cave paintings and remnants of Spanish colonization at Mission Santa Barbara. To this day, Catholic and Buddhist monasteries can be found dotted up the coast of Central California thanks to the region’s natural beauty and isolation.
We hope you find inspiration in the incredible plant life of this region. Be sure to check out the Coastal California: Monterey to Santa Barbara expedition!
Here are just a few native plant species featured in our expedition:
Eriastrum densifolium:
Eriastrum densifolium, also commonly known as the Giant Woollystar, is a member of the Phlox genus. This flowering plant is native California and Baja California, and grows primarily in sand dunes or dry washes.
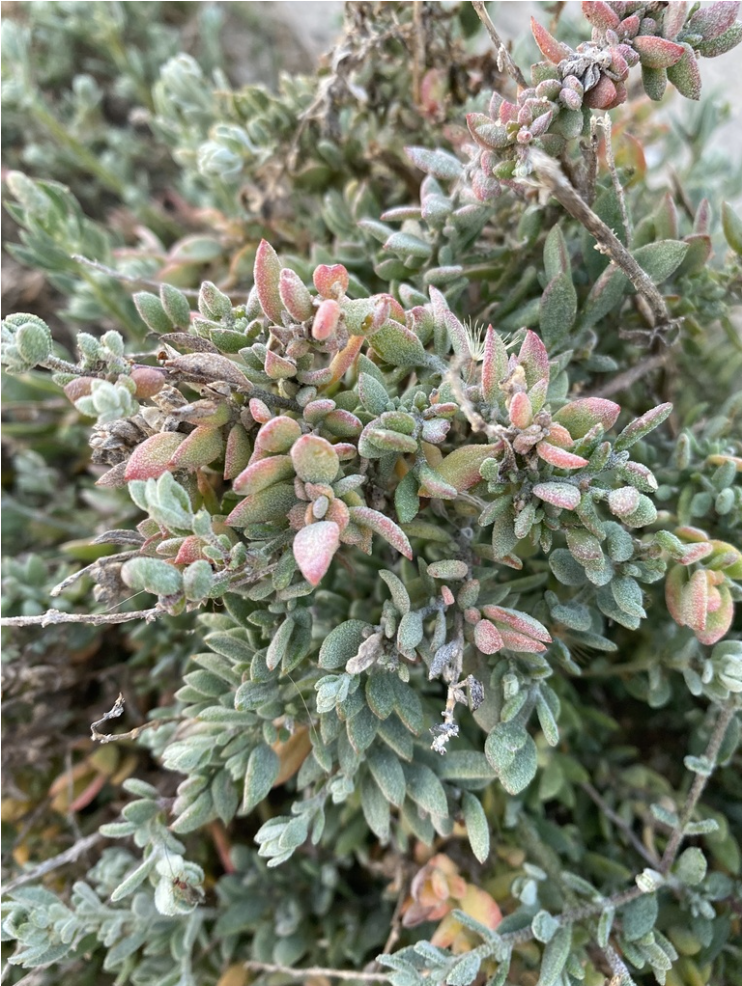
Extriplex Californica:
Exptriplex Californica is also known as California Saltbush and resides in coastal Salt marsh habitats of the California and Baja California coasts. Sometimes members of the Extriplex genus are called “Orach”, a name given to them in Latin by roman polymath Pliny the Elder.
Zeltnera muehlenbergii:
Zeltnera muehlenbergii is also known as the “Monterey centaury” but can also be found in other regions along the coast. This plant can be found mostly in forests and moist environments. Unlike many plants that flower in the Spring, this species flowers in the Summer and early Fall.