FAQs and Useful Tools
We have put together a list of frequently asked questions and useful tools related to transcribing records. These are based on discussion threads in the NfN discussion forums (linked at the bottom). Please take a look, and let us know if there are any questions we missed or useful tools that you can suggest. These will eventually go on a page of the Notes from Nature website.
NOTE: The herbarium interface (SERNEC), now has it’s own FAQ. It can be found here: https://blog.notesfromnature.org/2015/04/14/updated-faq-and-useful-tools-herbarium-interface/
We really appreciate your input and all that you do for Notes from Nature. Thank You!
Common issues or questions that you may encounter while transcribing Notes from Nature records:
1.) Interpretation: In general, you should minimize interpretation of open-ended fields and enter information verbatim. This way, we can better achieve consensus when checking multiple records against one another (see below, on that process). However, some discretion would be nice. Here are examples:
Interpretation that you should make: Simple spacing errors (e.g. “3miN. of Oakland” should be “3 mi N. of Oakland”)
Interpretation you should leave to us: Don’t interpret abbreviations, we’ll sort that out. (e.g. “Convict Lk.” )
2.) Not in English: Transcribe exactly as written. Match label content to transcription fields as best as you can.
3.) Abbreviations: Transcribe exactly as written.
4.) Spelling mistakes: Transcribe exactly as written, unless you have looked it up and are absolutely certain of a simple spelling mistake. In this case, you can enter the correct spelling.
5.) Problem records: If you come across a problem record that may need to be addressed by a scientist, like a faulty image or a record with illegible handwriting, you can flag the record by commenting on it (e.g. with the hashtag #error) and indicate what is in error.
6.) Provinces: Provinces go in the Location field (e.g. Coastal Plain Province, Piedmont Province).
7.) Capitalization: Sometimes information may be in all capital letters on the labels. Unless this is an abbreviation, you should capitalize only the first letter of every word in your transcription (e.g. “COASTAL PLAIN PROVINCE” should be “Coastal Plain Province”).
8.) Many collectors: In many cases, collectors may be listed on different lines of the label with no punctuation separating them. In your transcription, please separate the collectors with commas.
9.) Missing information: What should you do when there is no information available for a field? When information is not given on the label, you should leave the field blank (in the case of open-ended fields) or select “Unknown” or “Not Shown” in the drop-down lists.
10.) Inconsistent collector names: You will often find several variations of the same collector name (e.g. “R. Kral” or “R.Kral”, “RWG” or “R.W.Garrison”). Use similar discretion when transcribing these variations as in the localities.
Interpretation that you should make: Simple spacing errors (e.g. “R.Kral” should be “R. Kral”)
Interpretation you should leave to us: Don’t interpret abbreviations, we’ll sort that out. (e.g. “RWG” should remain “RWG”)
11.) Many scientific names: For Calbug, you do not need to enter the species name (we have this info already), but if there is a scientific name that is different from what is listed in the record, put it into the “Other Notes” field. These could be old names, or plant host names.
For SERNEC Herbarium specimens, copy only the most recent name. This can be determined based on the date that appears on the ‘annotation label.’ If you do not see a date then enter the name that appears on the primary label.
12.) Variations and subspecies (SERNEC): Record the subspecies, but omit the scientific author’s name. So “Cyperus odoratus var. squarrosus (Britton) Jones, Wipff & Carter” becomes “Cyperus odoratus var. squarrosus”. “Echinodorus cordifolius (Linnaeus) Grisebach ssp. cordifolius” becomes “Echinodorus cordifolius ssp. cordifolius”.
13.) Scientific name (SERNEC): Provide the most recent name, whether it is a species name (a two-word combination of the genus and what is called the “specific epithet” in botanical nomenclature) or a one-word name that is at a higher taxonomic rank (e.g., just the genus or family name). Names at higher taxonomic ranks than species are used when a more precise identification has not been made. The species name should typically take the form of a genus name that begins with a capital letter and a specific epithet that begins with a lowercase letter. If any of the names are given in all capitals, such as “CYPERUS ODORATUS”, the name should be entered using the typical convention, “Cyperus odoratus” in this case.
14.) Latitude and Longitude: How do you enter latitude and longitude values, and where do these values go? Enter exactly as written, you can find symbols in Word or by searching online (e.g. 33° 62’ 22” N 116° 41’ 42” W). You can also produce the degree symbol ° using key combinations (alt + 0 on a mac; alt + 0176 on a PC, with the key pad on the right side of your keyboard). This information should go into the “Location” or “Locality” field, depending on the project you are working on.
15.) Special Characters: What should you type when there is a special character in a text string, such as a degree symbol or language-specific characters? You can do a google search for the symbol or copy and paste it from Microsoft Word symbols. There are also key combinations for common symbols. As mentioned above, you can produce the degree symbol ° using key combinations (alt + 0 on a mac; alt + 0176 on a PC, with the key pad on the right side of your keyboard).
16.) Elevation: Enter elevation verbatim into the “Other Notes” field for Calbug and the “Habitat and Description” field for SERNEC Herbarium records.
17.) County: If the county is not given on the label, please find the appropriate county using google search. However, if there are multiple potential counties for a locality, please leave the county field blank.
18.) Checking your transcription: You can use the link to the left of the “Finish Record” button (e.g. “1/9” or “9/9”) to check the information that you entered. Just click on any of the fields to make any necessary edits to your transcription.
19.) When is a record finished?: These blog posts describe the data checking process that uses 4 transcriptions of the same record to derive a consensus.
https://blog.notesfromnature.org/2014/01/14/checking-notes-from-nature-data/
http://soyouthinkyoucandigitize.wordpress.com/2014/01/14/412/
Some Useful Tools (discovered by NfN users)
Counties and Cities: Good tools for finding counties etc. are lists on wikipedia, there are lists of municipalities in each state of the USA (there are also similar lists for others). For example, https://en.wikipedia.org/wiki/List_of_municipalities_in_Florida (via the linkbox you can also change the state).
Mountains: https://en.wikipedia.org/wiki/Category:Lists_of_mountains_of_the_United_States
Uncertain Localities: Geographic Names Information System, U.S. Geological Survey.
https://geonames.usgs.gov/pls/gnispublic
Mapping tool with topo quads: To find uncertain counties or localitieshttp://mapper.acme.com
Collector Names: The Essig Museum of Entomology database has lists of collector names and periods of activity for many collectors you will find in Calbug records. http://essigdb.berkeley.edu/query_people.html
Hard-to-read text: Use “Sheen”, the visual webpage filter, for some hard-to-read handwriting written in pencil. (Tip was from the War Diary Zooniverse project) https://chrome.google.com/webstore/detail/sheen/mopkplcglehjfbedbngcglkmajhflnjk?hl=en-GB
Special symbols: You should be able to find symbols in word or by doing a google search and copy and paste. Here are a few:
– degree symbol for coordinates: °
– plus minus: ±
– fractions: ⅛ ¼ ⅓ ⅜ ½ ⅝ ⅔ ¾ ⅞
– non-English symbols: Ä ä å Å ð ë ğ Ñ ñ õ Ö ö Ü ü Ž ž
The Plant List: Search for scientific names of plants –http://www.theplantlist.org/
List of Trees: https://en.wikipedia.org/wiki/Category:Trees_of_the_United_States
Integrated Taxonomic Information System (ITIS): http://www.itis.gov/
Essig database (for Calbug): Unsure if you spelled a collector name or species name right? It might also be worth checking similar entries already in the database, http://essigdb.berkeley.edu/.
NfN Discussion threads that this is based on:
FAQs
http://talk.notesfromnature.org/#/boards/BNN0000001/discussions/DNN000024q
Useful Tools
http://talk.notesfromnature.org/#/boards/BNN0000001/discussions/DNN00001vl
An Ode to Collecting: following the path of an early 20th century dragonfly collector
As you enter information from old labels, have you ever wondered about the collector’s experience—20, 60, or even 100 years ago? Have you thought about what the river, lake or meadow was like? Or how the place may have changed since the collector was there? I have, and so I spent a couple of years revisiting sites originally sampled by C.H. Kennedy for dragonflies in 1914-1915. Check out my recent blog post on this experience, entitled An ode to collecting: following the path of an early 20th century dragonfly collector.
Checking Notes from Nature Data
You may have wondered how we will use the data you are transcribing… Especially after you spend a few minutes trying to figure out the meaning of some unclear handwriting, and are uncertain whether the year on the label actually says “1976” or “1879,” or if the locality is “Vino” or “Vina” in California.
The first step in using Notes from Nature transcription data in research is checking data quality. Each specimen is transcribed independently by four different users, so mistakes in a single transcription do not impact the accuracy of our data very much. But how do we determine the right one out of the multiple transcriptions? For controlled fields such as country and state, this is simple: we perform a “vote-counting” procedure, where the value with the most “votes” is considered correct. However, this is much more difficult for open-ended fields such as locality or collector names.
One of the early approaches was to quantify differences between transcriptions. We did this by using the Levenshtein edit distance, named for the Russian information scientist, Vladimir Levenshtein, who developed a way to quantify the minimum number of single-character edits required to change one string of text to another. For example, “12 mi W Oakland” and “12 mi. W Oakland” would have a score of 1 because of the period after “mi”. We then looked for highly similar pairs of transcriptions for each specimen (pairs that were separated by the smallest edit distance), and then chose one of the pair. Our assumption was that if two separate transcriptions are almost identical to one another then either one must be very close to being correct. However, this assumption becomes violated if users are consistently wrong (either because the label is partially obscured, or the handwriting difficult to read, etc.). Furthermore, this was not a true consensus in the sense that our algorithm only decides which transcription out of the four is the best.
During the Hackathon last month (discussed in an earlier blog), it was quickly obvious that this approach was not ideal. In discussions with other researchers and experts, we developed an approach to combine the best elements of each individual transcription into a locality string that was greater than the sum of its parts. We did this using a concept from bioinformatics known as “sequence alignment.”
Genetics researchers and molecular ecologists use sequence alignment to match up multiple DNA or amino acid sequences in studies of evolutionary history or DNA mutation. To evaluate alignment, algorithms look for particular strings of amino acids or nucleotides within multiple sequences, and match similar or identical regions. Even closely related sequences, however, are often not identical and have small differences due to substitutions, deletions or insertions. So, alignment algorithms use complicated scoring mechanisms to generate alignments requiring as little “arrangement” as possible. The goal is to balance potential insertions, deletions and substitutions in characters from one sequence to the next.
In Notes from Nature data assessment, we use this method and assume that differences between transcriptions in a particular location may be due to substitutions. Or, we introduce gaps so that two separate chunks of sequence might be better aligned. This is especially useful if there are certain chunks of text within an individual transcription that are not consistently shared among all other transcriptions.
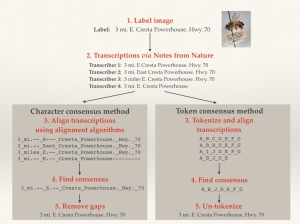
We implemented sequence alignment in two ways, using token and character alignment. For token alignment, we converted each chunk of text (usually individual words) and assigned each unique chunk a “token.” We then aligned the tokens. For character alignment, we compared each character in the text string directly. This proved to be the better method, as we inherently had a lot more information for the alignment software to work on.
An Illustration of Character and Token Alignment
To compare how well our algorithm was doing, we manually transcribed around 200 of Calbug’s specimens, and ran our algorithm on user transcriptions. 70% of the consensus transcriptions were either EXACTLY identical or within 5 edits to our manual transcriptions. While this result is preliminary and we’re still working on improvements, it is worth mentioning that the testing data set included quite a lot of the very worst examples of handwriting on a teeny tiny label.
Proportion of records that were exactly identical or within 5 edits to manual transcriptions
| Method |
Field |
Proportion exact (n =158) |
Proportion < 5 edits |
| Token |
Collector |
37.3 |
74.1 |
| Token |
Locality |
38.6 |
58.8 |
| Character |
Collector |
38.0 |
88.0 |
| Character |
Locality |
37.3 |
69.6 |
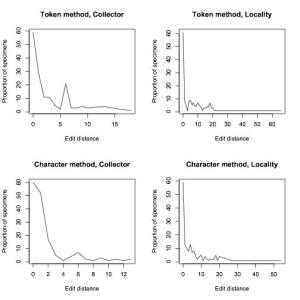
Proportion of specimens that were a certain edit distance away from our manually-transcribed set of specimens
In closing, we’d like to emphasize the importance of typing exactly what you see in the open-ended fields for collector and locality. A certain degree of interpretation may be necessary. For example, if there are no apparent spaces between words (e.g. DeathCanyonValley) it would be useful to add them. But, it is often very difficult (probably impossible) to derive a consensus if the inconsistency of label data is compounded by inconsistencies in data entry. We have ways of dealing with the huge diversity of abbreviations, so it would be helpful if you enter them as they are when you see them with minimal interpretation. The overall lesson from our data quality comparisons is that so far we are doing great, thanks to your careful work!
Want to learn more? Check out a related blog post, here.
—
By Junying Lim
Adventures in the field: How do insect museums get these specimens anyway?
As you pour over images of our fascinating CalBug specimens, you may ask yourself how these insects ended up in the museum in the first place. Many of the labels you are transcribing date back to 60-100 years ago, but don’t let that fool you into thinking that museums are places to that just store old specimens. Scientists are still adding to museum collections every day, but how we use specimens now is often in ways that Entomologists 60 years ago could not have imagined.
As a PhD student in the Essig Museum of Entomology, I have had many opportunities to work with insect specimens within a museum. However, this summer I had the chance to go on a month-long expedition in the Appalachian Mountains of North America to collect live insects in the field. My dissertation research involves understanding the diversification and evolution of ground beetles in the genus Scaphinotus. Often referred to as “snail-eaters,” these nocturnal beetles have developed an elongate head and mouthparts, including escargot fork-like jaws and huge sensory palps that allow them to find and feed on snails and slugs. They are flightless and live in predominantly montane habitats. This makes them interesting candidates for studying how body-forms of species change over time, possibly adapting to feeding preferences.
Insect specimens already housed in museums provide a great deal of information about morphology, distribution, seasonality and even behavior, however there is one thing they generally cannot provide- good quality DNA! So today entomologists are frequently heading to the field to collect specimens specifically to extract their DNA. This is why I went on my recent trip to the Appalachians, where I hoped to collect as many as 20 Scaphinotus species to use in my research.
A month-long field excursion requires careful planning and preparation. My trip included visits to 5 states and as many National Forests, where I camped and hiked long-forgotten trails in search of these elusive little beetles. Of course no amount of planning can prevent one from running into a month-long bout of stormy weather! And so it was, my first big trip into the field was vexed by torrential rains, flooding, lightning, thunder, and even a tornado! But in spite of all that heavy weather, rain and mud, I did manage to find a few Scaphinotus (the beetles were possibly as unhappy about the weather as I was!).
I came away from the trip with a far greater understanding and appreciation of what it is like to be in the field collecting specimens first hand. I also chalked up nine additional species whose DNA will contribute to my dissertation research, and will be made available to other scientists worldwide via CalBug and the Essig Museum of Entomology at UC Berkeley.
-Meghan Culpepper
Behind the scenes at the CalBug Project.
http://www.calacademy.org/sciencetoday/calbug/5513353/
Wonder where all those specimen images come from? Who are the people working in the museums? What does it look like? Take a behind the scenes look at the Essig Museum and the CalBug project in a new video produced by the California Academy of Sciences.